Hi-CO unravels the complex packing of nucleosomes
High-resolution genome structural analyses combined with large-scale simulations show the arrangements of the genome’s spool-like structures affecting gene expression.
Scientists at Kyoto University’s Institute for Integrated Cell-Material Sciences (iCeMS) in Japan have developed a technology that produces high-resolution simulations of one of the basic units of our genomes, called the nucleosome. Their findings were published in the journal Nature Protocols and should help improve understanding of how changes in nucleosome folding influence the inner workings of genes.
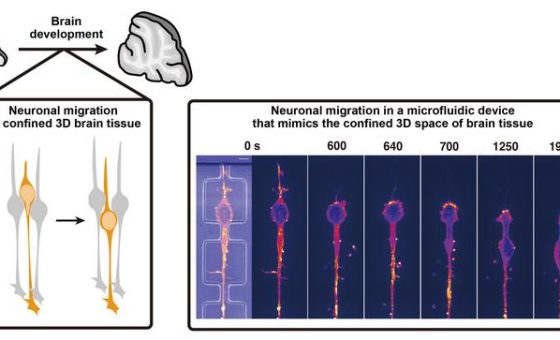
Nucleosomes are the basic structural units of DNA packaging inside the nucleus. They are formed of DNA wrapped around a small number of histone proteins. Nucleosomes move around inside the nucleus, folding and unfolding, changing their orientations, and moving closer together or further apart. These movements affect the accessibility of various molecules to DNA, determining when and how genes turn on and off.
“The 3D genome structure provides the physical molecular basis of gene expression processes in cells,” explains iCeMS systems biologist Yuichi Taniguchi, who led the research.
To better visualize this structure, Taniguchi and colleagues developed Hi-CO, short for high-throughput chromosome conformation capture with nucleosome orientation. It builds on existing Hi-C technology by significantly improving resolution so that simulations show the 3D positions and orientations of every nucleosome analysed in a sample.
“Being able to analyse this structure should help clarify the origins and control principles of many biological phenomena, including cell differentiation and immunity,” says molecular biologist Masae Ohno, who conducted the experiments and analyses.
Hi-CO involves a month-long process in which enzymes and a variety of other molecules are used to treat an organism’s genome, ultimately breaking down its DNA into millions of fragments that were close to each other due to nucleosome proximity. These fragments are then sequenced and the data is entered into a simulation program that shows the most likely orientations of each nucleosome.
Taniguchi and his team successfully tested Hi-CO on a yeast genome. They aim to next use it to analyse the genomes of other organisms, while continuing to improve the technology. They also hope to use Hi-CO to study genome structures in various cell differentiation states and diseases.
Paper Information
"Hi-CO: 3D genome structure analysis with nucleosome resolution"
Authors: Masae Ohno, Tadashi Ando, David G Priest and Yuichi Taniguchi